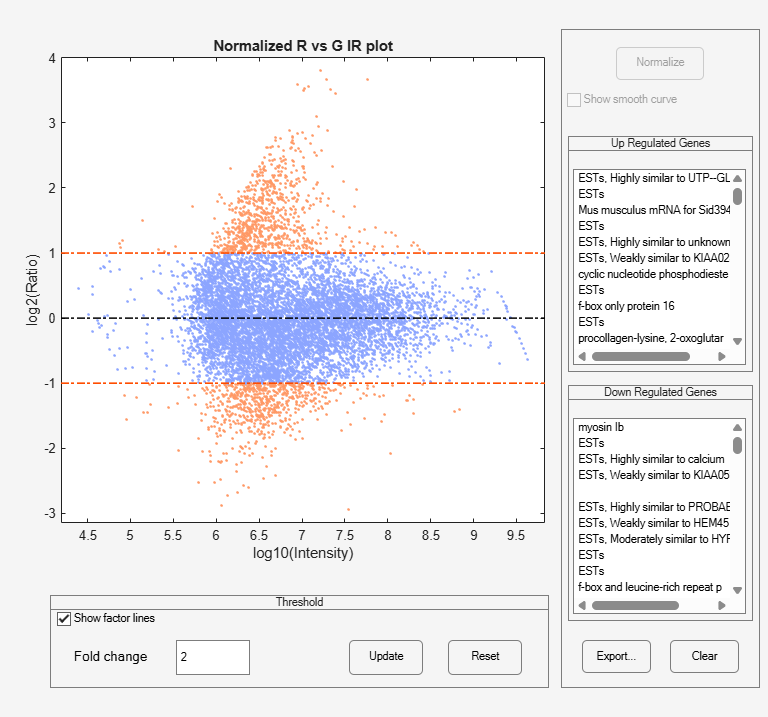
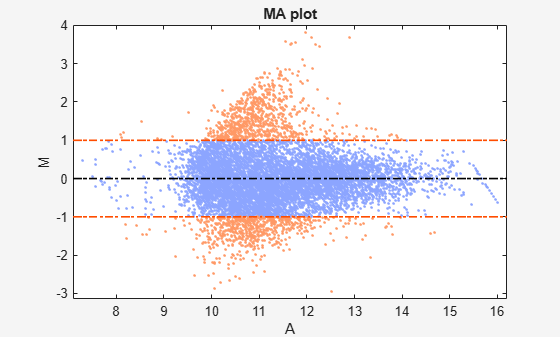
mairplot
Create intensity versus ratio scatter plot of microarray data
Syntax
Description
___
= mairplot(
specifies additional options using one or more name-value arguments. Use any arguments from
the previous syntaxes.DataX,DataY,Name=Value)
Examples
Input Arguments
Name-Value Arguments
Output Arguments
References
[1] Quackenbush, J. Microarray Data Normalization and Transformation. Nature Genetics suppl. 32 (2002): 496–501.
[2] Dudoit, S., Y.H. Yang, M.J. Callow, and T.P. Speed. Statistical Methods for Identifying Differentially Expressed Genes in Replicated cDNA Microarray Experiments. Statistica Sinica 12 (2002): 111–139.
Version History
Introduced before R2006a
See Also
DataMatrix | maboxplot | magetfield | maimage | mainvarsetnorm | maloglog | malowess | manorm | mattest | mavolcanoplot