summarize
Summarize threshold-switching dynamic regression model estimation results
Since R2021b
Description
summarize( displays a summary of the
threshold-switching dynamic regression model Mdl)Mdl.
If
Mdlis an estimated model returned byestimate, thensummarizedisplays estimation results to the MATLAB® Command Window. The display includes:A model description
Estimated threshold transitions
Fit statistics, which include the effective sample size, number of estimated submodel parameters and constraints, loglikelihood, and information criteria (AIC and BIC)
A table of submodel estimates and inferences, which includes coefficient estimates with standard errors, t-statistics, and p-values
If
Mdlis an unestimated threshold-switching model returned bytsVAR,summarizeprints the standard object display (the same display thattsVARprints during model creation).
results = summarize(___)
If
Mdlis an estimated threshold-switching model,resultsis a table containing the submodel estimates and inferences.If
Mdlis an unestimated model,resultsis atsVARobject that is equal toMdl.
summarize does not print to the Command Window
Examples
Input Arguments
Output Arguments
Algorithms
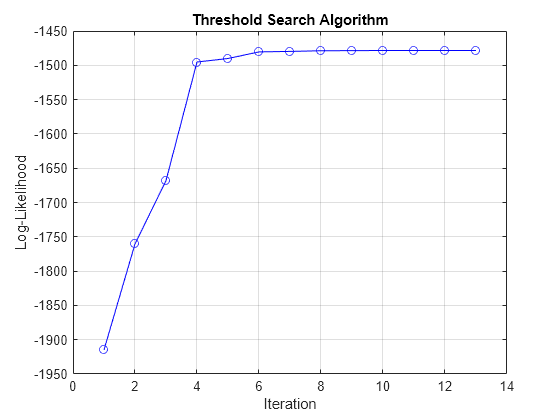
estimate searches over levels
and rates for estimated threshold transitions while solving a conditional least-squares
problem for submodel parameters, as described in [2]. The standard errors, loglikelihood, and
information criteria are conditional on optimal parameter values in the estimated threshold
transitions Mdl.Switch. In particular, standard errors do not account for
variation in estimated levels and rates.
References
Version History
Introduced in R2021b