Motor Current Signature Analysis for Gear Train Fault Detection
This example shows how to identify faults in a gear train using motor current signature analysis (MCSA) of a current signal driving a hobby-grade servo. MCSA is a useful method for the diagnosis of faults that induce torque or speed fluctuations, and has been proven to be ideal for motor fault analysis. Gear fault detection using traditional vibration instruments is challenging, especially in cases where the gear train is not easily accessible for instrumentation with accelerometers or other vibration sensors, like the inner workings of a nuclear power plant. This example illustrates how to apply current signature analysis to extract spectral metrics to detect faults in drive gears of a hobby-grade servo motor. The simplified workflow to obtain the spectral metrics from the current signal is as follows:
Compute nominal rpm to detect frequencies of interest.
Construct frequency bands where fault signals may be present.
Extract power spectral density (PSD) data.
Compute spectral metrics at the frequency bands of interest.
Hardware Overview

For this example, the electrical current data was collected from a standard Futaba S3003 hobby servo, which was modified for continuous rotation. Servos convert the high speed of the internal DC motor to high torque at the output spline. To achieve this, servos consist of a DC motor, a set of nylon or metal drive gears, and the control circuit. The control circuit was removed to allow the current signal to the DC motor to be directly monitored. The tachometer signal at the output spline of the servo was collected using an infrared photointerrupter along with a 35 mm diameter, eight-holed, standard hobby servo wheel. The eight holes in the wheel were equally spaced and the IR photointerrupter was placed such that there were exactly eight pulses per rotation of the servo wheel horn. The servo and photointerrupter were held in place by custom 3-D printed mounts.

The DC motor was driven at a constant 5 volts, and with four pairs of gears providing 278:1 speed reduction, the shaft speed at the spline was about 50 rpm. The current consumption was calculated using Ohm's law by measuring the voltage drop across a 0.5 Ohm resistor. Since the change in current measurement values was too small for detection, the current signal was amplified using a single-supply sensor interface amplifier. The amplified current signal was then filtered using an anti-aliasing fifth-order elliptic low-pass filter to smooth it and to eliminate noise before sending it to an Arduino® Uno through an analog-to-digital converter (ADC).

As the flow chart shows, the current signal was first amplified and filtered using the amplifier and the anti-aliasing low-pass filter, respectively. The Arduino Uno sampled the current signal through an ADC at 1.5 kHz and streamed it to the computer along with the tachometer pulses as serial data at a baud rate of 115200. A MATLAB® script fetched the serial data from the Arduino Uno and wrote it to a comma-separated values (CSV) file. The CSV files were then read and processed using MATLAB to extract the spectral metrics.
Servo Gear Train

The Futaba S3003 servo consists of four pairs of nylon gears as illustrated in the above figure. The pinion P1 on the DC motor shaft meshes with the stepped gear G1. The pinion P2 is a molded part of the stepped gear G1 and meshes with the stepped gear G2. The pinion P3, which is a molded part of gear G2, meshes with the stepped gear G3. Pinion P4, which is molded with G3, meshes with the final gear G4 that is attached to the output spline. The stepped gear sets G1 and P2, G2 and P3, and G3 and P4 are free spinning gears that is, they are not attached to their respective shafts. The set of drive gears provides a 278:1 reduction going from a motor speed of 13901 rpm to about 50 rpm at the output spline when the motor is driven at 5 volts. The following table outlines the tooth count and theoretical values of output speed, gear mesh frequencies, and cumulative gear reduction at each gear mesh.

A total of 10 healthy data sets were collected before the faults were introduced in the stepped gears G2 and G3. Since the gears were molded out of nylon, simulated cracks were introduced in both the gears by cutting slots in the tooth space with a hobby knife. The tooth space is the gap between two adjacent teeth measured along the pitch circle of the spur gear. The slot depths were about 70 percent of the gear radius. A total of 10 faulty data sets were recorded after introducing the faults in the gears G2 and G3.

Visualize Data
The file mcsaData.mat contains servoData, a 10-by-2 table of timetables where each timetable corresponds to one data set. The first column of servoData contains 10 timetables of healthy data while the second column contains 10 timetables of faulty data. Each data set contains about 11 seconds of data sampled at 1500 Hz.
Load the data.
load('mcsaData.mat','servoData') servoData
servoData=10×2 table
healthyData faultyData
___________________ ___________________
{16384×2 timetable} {16384×2 timetable}
{16384×2 timetable} {16384×2 timetable}
{16384×2 timetable} {16384×2 timetable}
{16384×2 timetable} {16384×2 timetable}
{16384×2 timetable} {16384×2 timetable}
{16384×2 timetable} {16384×2 timetable}
{16384×2 timetable} {16384×2 timetable}
{16384×2 timetable} {16384×2 timetable}
{16384×2 timetable} {16384×2 timetable}
{16384×2 timetable} {16384×2 timetable}
head(servoData.healthyData{1,1}) Time Pulse Signal
______________ _____ ______
0 sec 0 66.523
0.00066667 sec 0 62.798
0.0013333 sec 0 63.596
0.002 sec 0 64.128
0.0026667 sec 0 60.669
0.0033333 sec 0 62.798
0.004 sec 0 65.459
0.0046667 sec 0 56.678
Each timetable in servoData contains one data set with the tachometer signal in the first column and the current signal in the second column.
For this example, consider one set each of healthy and faulty data, and visualize the current signal and tachometer pulses.
healthyData = servoData.healthyData{1,1};
faultyData = servoData.faultyData{1,1};
healthyCurrent = healthyData.Signal;
faultyCurrent = faultyData.Signal;
healthyTacho = healthyData.Pulse;
faultyTacho = faultyData.Pulse;
tHealthy = healthyData.Time;
tFaulty = faultyData.Time;
figure
ax1 = subplot(221);
plot(tHealthy,healthyCurrent)
title('Current Signal - Healthy Gears')
ylabel('Current (mA)')
ax2 = subplot(222);
plot(tFaulty,faultyCurrent)
title('Current Signal - Faulty Gears')
ylabel('Current (mA)')
ax3 = subplot(223);
plot(tHealthy,healthyTacho)
title('Tachometer Pulse - Healthy Gears')
ylabel('Pulses, 8/rev')
ax4 = subplot(224);
plot(tFaulty,faultyTacho)
title('Tachometer Pulse - Faulty Gears')
ylabel('Pulses, 8/rev')
linkaxes([ax1,ax2,ax3,ax4],'x');
ax1.XLim = seconds([0 2]);
The output spline of the servo has eight pulses per rotation, corresponding to the eight holes on the servo wheel.
Compute Nominal Rpm
Compute the nominal speed to detect frequencies of interest in the gear system and match them correctly with the frequencies on the power spectra. Using the sampling frequency value of 1500 Hz, visualize the tachometer signal and compute the nominal rpm using tachorpm.
fs = 1500;
figure
tachorpm(healthyTacho,fs,'PulsesPerRev',8)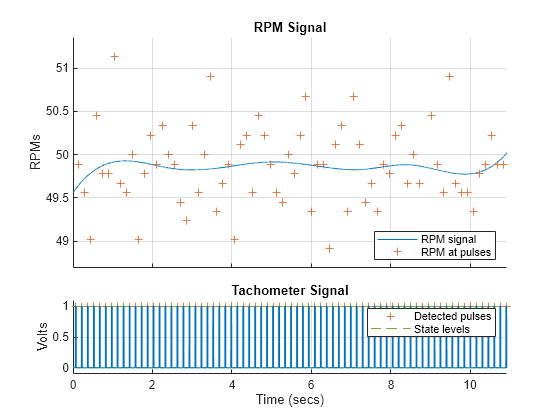
figure
tachorpm(faultyTacho,fs,'PulsesPerRev',8)
rpmHealthy = mean(tachorpm(healthyTacho,fs,'PulsesPerRev',8))rpmHealthy = 49.8550
rpmFaulty = mean(tachorpm(faultyTacho,fs,'PulsesPerRev',8))rpmFaulty = 49.5267
Observe that there is a very small difference in the output shaft speed between the healthy and faulty data sets. The actual nominal rpm values are also close to the theoretical value of 50 rpm. Hence, consider the same value of 49.9 rpm for both the healthy and faulty signal analysis.
Construct Frequency Bands
Constructing frequency bands is an important prerequisite for computing the spectral metrics. Using the tooth count of the drive gears in the gear train and the nominal rpm, first compute the frequencies of interest. The frequencies of interest are actual output speed values in hertz whose values are close to the theoretical values listed in the table below.

G4 = 41; G3 = 35; G2 = 50; G1 = 62; P4 = 16; P3 = 10; P2 = 10; P1 = 10; rpm = rpmHealthy; FS5 = mean(rpm)/60; FS4 = G4/P4*FS5; FS3 = G3/P3*FS4; FS2 = G2/P2*FS3; FS1 = G1/P1*FS2; FS = [FS1,FS2,FS3,FS4,FS5]
FS = 1×5
231.0207 37.2614 7.4523 2.1292 0.8309
Next, construct the frequency bands for the all the output speeds which include the following frequencies of interest using faultBands.
FS1 at 231 Hz, its first two harmonics, and 0:1 sidebands of FS2
FS2 at 37.3 Hz, its first two harmonics, and 0:1 sidebands of FS3
FS3 at 7.5 Hz and its first two harmonics
FS4 at 2.1 Hz and its first two harmonics
[FB1,info1] = faultBands(FS1,1:2,FS2,0:1); [FB2,info2] = faultBands(FS2,1:2,FS3,0:1); [FB3,info3] = faultBands(FS3,1:2); [FB4,info4] = faultBands(FS4,1:2); FB = [FB1;FB2;FB3;FB4]
FB = 16×2
187.9838 199.5348
225.2452 236.7962
262.5066 274.0577
419.0045 430.5556
456.2659 467.8170
493.5273 505.0784
28.8776 30.7407
36.3299 38.1929
43.7822 45.6452
66.1390 68.0021
73.5913 75.4543
81.0436 82.9066
7.2660 7.6386
14.7183 15.0909
2.0760 2.1825
⋮
The output FB is a 16-by-2 array of frequency ranges for these frequencies of interest.
Extract Power Spectral Density (PSD) Data
Compute and visualize the power spectrum of the healthy and faulty data. Also plot the frequencies of interest on the spectrum plot by using the information in the structure info.
figure pspectrum(healthyCurrent,fs,'FrequencyResolution',0.5) hold on pspectrum(faultyCurrent,fs,'FrequencyResolution',0.5) hold on info1.Labels = regexprep(info1.Labels,'F0','FS1'); info1.Labels = regexprep(info1.Labels,'F1','FS2'); helperPlotXLines(info1,[0.5 0.5 0.5]) info2.Labels = regexprep(info2.Labels,'F0','FS2'); info2.Labels = regexprep(info2.Labels,'F1','FS3'); helperPlotXLines(info2,[0.5 0.5 0.5]) info3.Labels = regexprep(info3.Labels,'F0','FS3'); helperPlotXLines(info3,[0.5 0.5 0.5]) info4.Labels = regexprep(info4.Labels,'F0','FS4'); helperPlotXLines(info4,[0.5 0.5 0.5]) legend('Healthy Data','Faulty Data','Location','South') hold off

The plot in blue indicates the healthy spectrum while the plot in red indicates the spectrum of the faulty data. From the plot, observe the increase in amplitudes of fault frequencies:
1FS1 at 231 Hz, its second harmonic 2FS1 at 462 Hz, and their respective sidebands
Zoom in on the plot to observe the increase in amplitudes of the following remaining frequencies:
1FS2 at 37.2 Hz and its sidebands
1FS3 at 7.5 Hz and its second harmonic 2FS3 at 15 Hz
1FS4 at 2.1 Hz and its second harmonic 2FS4 at 4.2 Hz

Use pspectrum to compute and store the PSD and the frequency grid data for the healthy and faulty signals respectively.
[psdHealthy,fHealthy] = pspectrum(healthyCurrent,fs,'FrequencyResolution',0.5); [psdFaulty,fFaulty] = pspectrum(faultyCurrent,fs,'FrequencyResolution',0.5);
Compute Spectral Metrics
Using the frequency bands and the PSD data, compute the spectral metrics for the healthy and faulty data using faultBandMetrics.
spectralMetrics = faultBandMetrics({[psdHealthy,fHealthy],[psdFaulty,fFaulty]},FB)spectralMetrics=2×49 table
PeakAmplitude1 PeakFrequency1 BandPower1 PeakAmplitude2 PeakFrequency2 BandPower2 PeakAmplitude3 PeakFrequency3 BandPower3 PeakAmplitude4 PeakFrequency4 BandPower4 PeakAmplitude5 PeakFrequency5 BandPower5 PeakAmplitude6 PeakFrequency6 BandPower6 PeakAmplitude7 PeakFrequency7 BandPower7 PeakAmplitude8 PeakFrequency8 BandPower8 PeakAmplitude9 PeakFrequency9 BandPower9 PeakAmplitude10 PeakFrequency10 BandPower10 PeakAmplitude11 PeakFrequency11 BandPower11 PeakAmplitude12 PeakFrequency12 BandPower12 PeakAmplitude13 PeakFrequency13 BandPower13 PeakAmplitude14 PeakFrequency14 BandPower14 PeakAmplitude15 PeakFrequency15 BandPower15 PeakAmplitude16 PeakFrequency16 BandPower16 TotalBandPower
______________ ______________ __________ ______________ ______________ __________ ______________ ______________ __________ ______________ ______________ __________ ______________ ______________ __________ ______________ ______________ __________ ______________ ______________ __________ ______________ ______________ __________ ______________ ______________ __________ _______________ _______________ ___________ _______________ _______________ ___________ _______________ _______________ ___________ _______________ _______________ ___________ _______________ _______________ ___________ _______________ _______________ ___________ _______________ _______________ ___________ ______________
0.0020712 193.75 0.010881 0.50813 231.06 0.46652 0.0019579 272.5 0.011841 0.0020486 424.06 0.011225 0.54982 462 0.89544 0.0024293 493.69 0.0091045 0.002966 29.812 0.0035485 0.015582 37.25 0.011132 0.0028865 44.688 0.0041317 0.011896 67.062 0.0072753 0.059126 74.5 0.033568 0.013218 82 0.0079886 5.7904 7.4375 2.3115 0.068452 14.938 0.027653 0.79006 2.125 0.14382 0.09849 4.25 0.01806 3.9737
0.009804 192.44 0.017916 3.6921 229.44 2.9975 0.0035204 266.44 0.015639 0.0057056 421.75 0.019293 1.2974 459.69 3.2185 0.0053261 495.88 0.016296 0.0031674 28.938 0.0044271 0.023983 37 0.014447 0.0136 44.438 0.0089119 0.011419 66.625 0.0077035 0.0684 74 0.037016 0.012244 81.438 0.0075805 7.7931 7.375 3.0193 0.15692 14.812 0.058255 2.4211 2.125 0.4407 0.55167 4.25 0.10029 9.9838
The output is a 2-by-49 table of metrics for the frequency ranges in FB. The first row contains the metrics for healthy data while the second row contains the faulty data metrics. Observe that the following metrics have considerably higher values for the faulty data than for the healthy data:
PeakAmplitude2for the frequency at 231 Hz with a difference of 3.1842 unitsTotalBandPowerwith a difference of 6.01 units
Hence, use these two metrics to create a scatter plot to group the faulty and healthy data sets separately.
Create Scatter Plot
Create a scatter plot to classify healthy and faulty data using the two spectral metrics PeakAmplitude2 and TotalBandPower for all 20 data sets in the table servoData.
plotData = zeros(10,4); for n = 1:max(size(servoData)) hC = servoData.healthyData{n,1}.Signal; fC = servoData.faultyData{n,1}.Signal; [psdH,freqH] = pspectrum(hC,fs,'FrequencyResolution',0.5); [psdF,freqF] = pspectrum(fC,fs,'FrequencyResolution',0.5); sMetrics = faultBandMetrics({[psdH,freqH],[psdF,freqF]},FB); plotData(n,:) = [sMetrics{:,4}',sMetrics{:,49}']; end figure scatter(plotData(:,1),plotData(:,3),[],'blue') hold on; scatter(plotData(:,2),plotData(:,4),[],'red') legend('Healthy Data','Faulty Data','Location','best') xlabel('Peak Amplitude 2') ylabel('Total Band Power') hold off
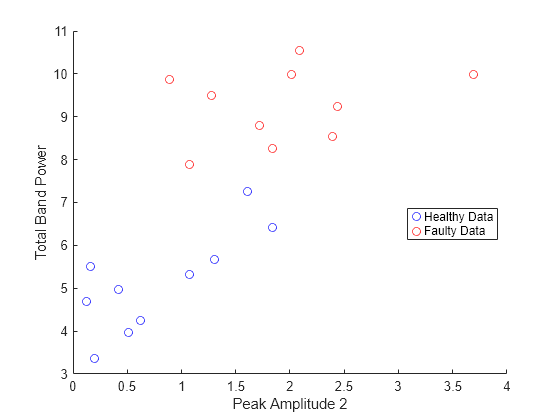
Observe that the healthy data sets and the faulty data sets are grouped in different areas of the scatter plot.
Hence, you can classify faulty and healthy data by analyzing the current signature of the servo motor.
To use all the available spectral metrics for classification, use Classification Learner.
Helper Function
The helper function helperPlotXLines uses the information in the structure info to plot the frequency band center lines on the power spectrum plot.
function helperPlotXLines(I,c) for k = 1:length(I.Centers) xline(I.Centers(k),'Label',I.Labels(k),'LineStyle','-.','Color',c); end end
References
[1] Moster, P.C. "Gear Fault Detection and Classification Using Learning Machines." Sound & vibration. 38. 22-27. 2004
See Also
Classification Learner | faultBands | faultBandMetrics | pspectrum