lsqnonlin
Solve nonlinear least-squares (nonlinear data-fitting) problems
Syntax
Description
Nonlinear least-squares solver
Solves nonlinear least-squares curve fitting problems of the form
subject to the constraints
x, lb, and ub can be vectors or
matrices; see Matrix Arguments.
Do not specify the objective function as the scalar value (the sum of squares). lsqnonlin requires the objective
function to be the vector-valued function
x = lsqnonlin(fun,x0)x0 and finds a minimum of the sum of squares of
the functions described in fun. The function fun
should return a vector (or array) of values and not the sum of squares of the values.
(The algorithm implicitly computes the sum of squares of the components of
fun(x).)
Note
Passing Extra Parameters explains how to pass extra parameters to the
vector function fun(x), if necessary.
x = lsqnonlin(fun,x0,lb,ub)x,
so that the solution is always in the range
lb ≤ x ≤ ub.
You can fix the solution component x(i) by specifying
lb(i) = ub(i).
Note
If the specified input bounds for a problem are inconsistent, the output
x is x0 and the outputs
resnorm and residual are
[].
Components of x0 that violate the bounds
lb ≤ x ≤ ub are reset to the interior of
the box defined by the bounds. Components that respect the bounds are not
changed.
Examples
Find the exponential decay rate for the nonlinear function
that best fits data. Generate the data from the expression
,
where goes from 0 through 3, and is normally distributed with a mean of 0 and standard deviation of 0.05.
rng default % For reproducibility t = linspace(0,3); y = exp(-1.3*t) + 0.05*randn(size(t));
Create an anonymous function that takes a value of the exponential decay rate and returns a vector of differences from the model with that decay rate and the data.
fun = @(r)exp(-r*t)-y;
Find the value of the optimal decay rate. Arbitrarily choose an initial guess x0 = 4.
x0 = 4; r = lsqnonlin(fun,x0)
Local minimum possible. lsqnonlin stopped because the final change in the sum of squares relative to its initial value is less than the value of the function tolerance. <stopping criteria details>
r = 1.2645
Plot the data and the best-fitting exponential curve.
plot(t,y,"o",t,exp(-r*t),"-") legend("Data","Best fit") xlabel("t") ylabel("exp(-r*t)")

Find the best-fitting model when some of the fitting parameters have bounds.
Find a centering parameter and scaling parameter that best fit the function
to data generated from the standard normal density
subject to the bound constraints
Create a vector t of data points, and the corresponding normal density at those points.
t = linspace(-4,4); y = 1/sqrt(2*pi)*exp(-t.^2/2);
Create a function that evaluates the difference between the centered and scaled function from the normal y, with x(1) as the scaling parameter and x(2) as the centering parameter .
fun = @(x)x(1)*exp(-t).*exp(-exp(-(t-x(2)))) - y;
Find the optimal fit starting from x0 = [1/2,0], with the scaling between 1/2 and 3/2, and the centering between –1 and 3.
lb = [1/2,-1]; ub = [3/2,3]; x0 = [1/2,0]; x = lsqnonlin(fun,x0,lb,ub)
Local minimum possible. lsqnonlin stopped because the final change in the sum of squares relative to its initial value is less than the value of the function tolerance. <stopping criteria details>
x = 1×2
0.8231 -0.2444
Plot the two functions to see the quality of the fit.
plot(t,y,"-",t,fun(x)+y,"-") xlabel("t") legend("Normal density","Fitted function")

Consider the following objective function, a sum of squares:
The code for this objective function appears as the myfun function at the end of this example.
Minimize this function subject to the linear constraint . Write this constraint as .
A = [1 -1/2]; b = 0;
Impose the bounds , , , and .
lb = [0 0]; ub = [2 4];
Start the optimization process from the point x0 = [0.3 0.4].
x0 = [0.3 0.4];
The problem has no linear equality constraints.
Aeq = []; beq = [];
Run the optimization.
x = lsqnonlin(@myfun,x0,lb,ub,A,b,Aeq,beq)
Local minimum found that satisfies the constraints. Optimization completed because the objective function is non-decreasing in feasible directions, to within the value of the optimality tolerance, and constraints are satisfied to within the value of the constraint tolerance. <stopping criteria details>
x = 1×2
0.1695 0.3389
function F = myfun(x) k = 1:10; F = 2 + 2*k - exp(k*x(1)) - 2*exp(2*k*(x(2)^2)); end
Consider the following objective function, a sum of squares:
The code for this objective function appears as the myfun function at the end of this example.
Minimize this function subject to the nonlinear constraint . The code for this nonlinear constraint function appears as the nlcon function at the end of this example.
Impose the bounds , , , and .
lb = [0 0]; ub = [2 4];
Start the optimization process from the point x0 = [0.3 0.4].
x0 = [0.3 0.4];
The problem has no linear constraints.
A = []; b = []; Aeq = []; beq = [];
Run the optimization.
x = lsqnonlin(@myfun,x0,lb,ub,A,b,Aeq,beq,@nlcon)
Local minimum found that satisfies the constraints. Optimization completed because the objective function is non-decreasing in feasible directions, to within the value of the optimality tolerance, and constraints are satisfied to within the value of the constraint tolerance. <stopping criteria details>
x = 1×2
0.2133 0.3266
function F = myfun(x) k = 1:10; F = 2 + 2*k - exp(k*x(1)) - 2*exp(2*k*(x(2)^2)); end function [ineqnonlin,eqnonlin] = nlcon(x) eqnonlin = []; ineqnonlin = sin(x(1)) - cos(x(2)); end
Compare the results of a data-fitting problem when using different lsqnonlin algorithms.
Suppose that you have observation time data xdata and observed response data ydata, and you want to find parameters and to fit a model of the form
Input the observation times and responses.
xdata = ... [0.9 1.5 13.8 19.8 24.1 28.2 35.2 60.3 74.6 81.3]; ydata = ... [455.2 428.6 124.1 67.3 43.2 28.1 13.1 -0.4 -1.3 -1.5];
Create a simple exponential decay model. The model computes a vector of differences between predicted values and observed values.
fun = @(x)x(1)*exp(x(2)*xdata)-ydata;
Fit the model using the starting point x0 = [100,-1]. First, use the default "trust-region-reflective" algorithm.
x0 = [100,-1];
options = optimoptions(@lsqnonlin,Algorithm="trust-region-reflective");
x = lsqnonlin(fun,x0,[],[],options)Local minimum possible. lsqnonlin stopped because the final change in the sum of squares relative to its initial value is less than the value of the function tolerance. <stopping criteria details>
x = 1×2
498.8309 -0.1013
See if there is any difference using the "levenberg-marquardt" algorithm.
options.Algorithm = "levenberg-marquardt";
x = lsqnonlin(fun,x0,[],[],options)Local minimum possible. lsqnonlin stopped because the relative size of the current step is less than the value of the step size tolerance. <stopping criteria details>
x = 1×2
498.8309 -0.1013
The two algorithms found the same solution. Plot the solution and the data.
plot(xdata,ydata,"ko") hold on tlist = linspace(xdata(1),xdata(end)); plot(tlist,x(1)*exp(x(2)*tlist),"b-") xlabel xdata ylabel ydata title("Exponential Fit to Data") legend("Data","Exponential Fit") hold off

Find the that minimizes
,
and find the value of the minimal sum of squares.
Because lsqnonlin assumes that the sum of squares is not explicitly formed in the user-defined function, the function passed to lsqnonlin should instead compute the vector-valued function
,
for to (that is, should have components).
The myfun function, which computes the 10-component vector F, appears at the end of this example.
Find the minimizing point and the minimum value, starting at the point x0 = [0.3,0.4].
x0 = [0.3,0.4]; [x,resnorm] = lsqnonlin(@myfun,x0)
Local minimum possible. lsqnonlin stopped because the size of the current step is less than the value of the step size tolerance. <stopping criteria details>
x = 1×2
0.2578 0.2578
resnorm = 124.3622
The resnorm output is the squared residual norm, or the sum of squares of the function values.
The following function computes the vector-valued objective function.
function F = myfun(x) k = 1:10; F = 2 + 2*k-exp(k*x(1))-exp(k*x(2)); end
Examine the solution process both as it occurs (by setting the Display option to "iter") and afterward (by examining the output structure).
Suppose that you have observation time data xdata and observed response data ydata, and you want to find parameters and to fit a model of the form
Input the observation times and responses.
xdata = ... [0.9 1.5 13.8 19.8 24.1 28.2 35.2 60.3 74.6 81.3]; ydata = ... [455.2 428.6 124.1 67.3 43.2 28.1 13.1 -0.4 -1.3 -1.5];
Create a simple exponential decay model. The model computes a vector of differences between predicted values and observed values.
fun = @(x)x(1)*exp(x(2)*xdata)-ydata;
Fit the model using the starting point x0 = [100,-1]. Examine the solution process by setting the Display option to "iter". Obtain an output structure to obtain more information about the solution process.
x0 = [100,-1]; options = optimoptions("lsqnonlin",Display="iter"); [x,resnorm,residual,exitflag,output] = lsqnonlin(fun,x0,[],[],options);
Norm of First-order
Iteration Func-count Resnorm step optimality
0 3 359677 2.88e+04
Objective function returned Inf; trying a new point...
1 6 359677 11.6976 2.88e+04
2 9 321395 0.5 4.97e+04
3 12 321395 1 4.97e+04
4 15 292253 0.25 7.06e+04
5 18 292253 0.5 7.06e+04
6 21 270350 0.125 1.15e+05
7 24 270350 0.25 1.15e+05
8 27 252777 0.0625 1.63e+05
9 30 252777 0.125 1.63e+05
10 33 243877 0.03125 7.48e+04
11 36 243660 0.0625 8.7e+04
12 39 243276 0.0625 2e+04
13 42 243174 0.0625 1.14e+04
14 45 242999 0.125 5.1e+03
15 48 242661 0.25 2.04e+03
16 51 241987 0.5 1.91e+03
17 54 240643 1 1.04e+03
18 57 237971 2 3.36e+03
19 60 232686 4 6.04e+03
20 63 222354 8 1.2e+04
21 66 202592 16 2.25e+04
22 69 166443 32 4.05e+04
23 72 106320 64 6.68e+04
24 75 28704.7 128 8.31e+04
25 78 89.7947 140.674 2.22e+04
26 81 9.57381 2.02599 684
27 84 9.50489 0.0619927 2.27
28 87 9.50489 0.000462262 0.0114
Local minimum possible.
lsqnonlin stopped because the final change in the sum of squares relative to
its initial value is less than the value of the function tolerance.
<stopping criteria details>
Examine the output structure to obtain more information about the solution process.
output
output = struct with fields:
firstorderopt: 0.0114
iterations: 28
funcCount: 87
cgiterations: 0
algorithm: 'trust-region-reflective'
stepsize: 4.6226e-04
message: 'Local minimum possible.↵↵lsqnonlin stopped because the final change in the sum of squares relative to ↵its initial value is less than the value of the function tolerance.↵↵<stopping criteria details>↵↵Optimization stopped because the relative sum of squares (r) is changing↵by less than options.FunctionTolerance = 1.000000e-06.'
bestfeasible: []
constrviolation: []
For comparison, set the Algorithm option to "levenberg-marquardt".
options.Algorithm = "levenberg-marquardt";
[x,resnorm,residual,exitflag,output] = lsqnonlin(fun,x0,[],[],options); First-order Norm of
Iteration Func-count Resnorm optimality Lambda step
0 3 359677 2.88e+04 0.01
Objective function returned Inf; trying a new point...
1 13 340761 3.91e+04 100000 0.280777
2 16 304661 5.97e+04 10000 0.373146
3 21 297292 6.55e+04 1e+06 0.0589933
4 24 288240 7.57e+04 100000 0.0645444
5 28 275407 1.01e+05 1e+06 0.0741266
6 31 249954 1.62e+05 100000 0.094571
7 36 245896 1.35e+05 1e+07 0.0133606
8 39 243846 7.26e+04 1e+06 0.00944311
9 42 243568 5.66e+04 100000 0.00821622
10 45 243424 1.61e+04 10000 0.00777936
11 48 243322 8.8e+03 1000 0.0673933
12 51 242408 5.1e+03 100 0.675209
13 54 233628 1.05e+04 10 6.59804
14 57 169089 8.51e+04 1 54.6992
15 60 30814.7 1.54e+05 0.1 196.939
16 63 147.496 8e+03 0.01 129.795
17 66 9.51503 117 0.001 9.96069
18 69 9.50489 0.0714 0.0001 0.0804859
19 72 9.50489 4.95e-05 1e-05 5.07024e-05
Local minimum possible.
lsqnonlin stopped because the relative size of the current step is less than
the value of the step size tolerance.
<stopping criteria details>
The "levenberg-marquardt" converged with fewer iterations, but almost as many function evaluations:
output
output = struct with fields:
iterations: 19
funcCount: 72
stepsize: 5.0702e-05
cgiterations: []
firstorderopt: 4.9454e-05
algorithm: 'levenberg-marquardt'
message: 'Local minimum possible.↵lsqnonlin stopped because the relative size of the current step is less than↵the value of the step size tolerance.↵↵<stopping criteria details>↵↵Optimization stopped because the relative norm of the current step, 1.016425e-07,↵is less than options.StepTolerance = 1.000000e-06.'
bestfeasible: []
constrviolation: []
Input Arguments
Function whose sum of squares is minimized, specified as a function handle or
the name of a function. For the 'interior-point' algorithm,
fun must be a function handle. fun is a
function that accepts an array x and returns an array
F, the objective function evaluated at
x.
Note
The sum of squares should not be formed explicitly. Instead, your function should return a vector of function values. See Examples.
The function fun can be specified as a function handle to a
file:
x = lsqnonlin(@myfun,x0)
where myfun is a MATLAB® function such as
function F = myfun(x) F = ... % Compute function values at x
fun can also be a function handle for an anonymous
function.
x = lsqnonlin(@(x)sin(x.*x),x0);
lsqnonlin passes x to your objective function in the shape of the x0 argument. For example, if x0 is a 5-by-3 array, then lsqnonlin passes x to fun as a 5-by-3 array.
If the Jacobian can also be computed and the
'SpecifyObjectiveGradient' option is
true, set by
options = optimoptions('lsqnonlin','SpecifyObjectiveGradient',true)
then the function fun must return a second output argument
with the Jacobian value J (a matrix) at x.
By checking the value of nargout, the function can avoid
computing J when fun is called with only one
output argument (in the case where the optimization algorithm only needs the value
of F but not J).
function [F,J] = myfun(x) F = ... % Objective function values at x if nargout > 1 % Two output arguments J = ... % Jacobian of the function evaluated at x end
If fun returns an array of m components
and x has n elements, where
n is the number of elements of x0, the
Jacobian J is an m-by-n
matrix where J(i,j) is the partial derivative of
F(i) with respect to x(j). (The Jacobian
J is the transpose of the gradient of
F.)
Example: @(x)cos(x).*exp(-x)
Data Types: char | function_handle | string
Initial point, specified as a real vector or real array. Solvers use the number of elements in
x0 and the size of x0 to determine the number
and size of variables that fun accepts.
Example: x0 = [1,2,3,4]
Data Types: double
Lower bounds, specified as a real vector or real array. If the number of elements in
x0 is equal to the number of elements in lb,
then lb specifies that
x(i) >= lb(i) for all i.
If numel(lb) < numel(x0), then lb specifies
that
x(i) >= lb(i) for 1 <=
i <= numel(lb).
If lb has fewer elements than x0, solvers issue a
warning.
Example: To specify that all x components are positive, use lb =
zeros(size(x0)).
Data Types: single | double
Upper bounds, specified as a real vector or real array. If the number of elements in
x0 is equal to the number of elements in ub,
then ub specifies that
x(i) <= ub(i) for all i.
If numel(ub) < numel(x0), then ub specifies
that
x(i) <= ub(i) for 1 <=
i <= numel(ub).
If ub has fewer elements than x0, solvers issue
a warning.
Example: To specify that all x components are less than 1, use ub =
ones(size(x0)).
Data Types: single | double
Linear inequality constraints, specified as a real matrix. A is an
M-by-N
matrix, where M is the number of
inequalities, and N is the number
of variables (number of elements in
x0). For large problems with
algorithms that support sparse data, pass
A as a sparse matrix. See Sparsity in Optimization Algorithms.
A encodes the M linear
inequalities
A*x <= b,
where x is the column vector of N variables x(:),
and b is a column vector with M elements.
For example, consider these inequalities:
x1 + 2x2 ≤
10
3x1 +
4x2 ≤ 20
5x1 +
6x2 ≤ 30,
Specify the inequalities by entering the following constraints.
A = [1,2;3,4;5,6]; b = [10;20;30];
Example: To specify that the x components sum to 1 or less, use A =
ones(1,N) and b = 1.
Data Types: single | double
Linear inequality constraints, specified as a real vector. b is an
M-element vector related to the A matrix. If
you pass b as a row vector, solvers internally convert
b to the column vector b(:).
b encodes the M linear
inequalities
A*x <= b,
where x is the column vector of N variables x(:),
and A is a matrix of size M-by-N.
For example, consider these inequalities:
x1
+ 2x2 ≤
10
3x1
+ 4x2 ≤
20
5x1
+ 6x2 ≤
30.
Specify the inequalities by entering the following constraints.
A = [1,2;3,4;5,6]; b = [10;20;30];
Example: To specify that the x components sum to 1 or less, use A =
ones(1,N) and b = 1.
Data Types: single | double
Linear equality constraints, specified as a real matrix. Aeq is an
Me-by-N
matrix, where Me is the number of
equalities, and N is the number
of variables (number of elements in
x0). For large problems with
algorithms that support sparse data, pass
A as a sparse matrix. See Sparsity in Optimization Algorithms.
Aeq encodes the Me linear
equalities
Aeq*x = beq,
where x is the column vector of N variables x(:),
and beq is a column vector with Me elements.
For example, consider these inequalities:
x1 + 2x2 +
3x3 = 10
2x1 +
4x2 + x3 =
20,
Specify the inequalities by entering the following constraints.
Aeq = [1,2,3;2,4,1]; beq = [10;20];
Example: To specify that the x components sum to 1, use Aeq = ones(1,N) and
beq = 1.
Data Types: single | double
Linear equality constraints, specified as a real vector. beq is an
Me-element vector related to the Aeq matrix.
If you pass beq as a row vector, solvers internally convert it to the
column vector beq(:).
beq encodes the Me linear
equalities
Aeq*x = beq,
where x is the column vector of N variables
x(:), and Aeq is a matrix of size
Me-by-N.
For example, consider these equalities:
x1
+ 2x2 +
3x3 =
10
2x1
+ 4x2 +
x3 =
20.
Specify the equalities by entering the following constraints.
Aeq = [1,2,3;2,4,1]; beq = [10;20];
Example: To specify that the x components sum to 1, use Aeq = ones(1,N) and
beq = 1.
Data Types: single | double
Nonlinear constraints, specified as a function handle or function name.
nonlcon is a function that accepts a vector or array
x and returns two arrays, ineqnonlin(x) and
eqnonlin(x).
ineqnonlin(x)is the array of nonlinear inequality constraints atx.lsqnonlinattempts to satisfyineqnonlin(x) <= 0for all entries ofineqnonlin.eqnonlin(x)is the array of nonlinear equality constraints atx.lsqnonlinattempts to satisfyeqnonlin(x) = 0for all entries ofeqnonlin.
Note
The returned size of a nonlinear constraint function must not change during a solver run.
For example,
x = lsqnonlin(@myfun,x0,...,@mycon)
where mycon is a MATLAB function such as the
following:
function [ineqnonlin,eqnonlin] = mycon(x) ineqnonlin = ... % Compute nonlinear inequalities at x. eqnonlin = ... % Compute nonlinear equalities at x.
Suppose that the gradients of the constraints can also be computed and
the SpecifyConstraintGradient option is true, as
set by:
options =
optimoptions("lsqnonlin",SpecifyConstraintGradient=true)
In this case, the function nonlcon must also return, in the third and
fourth output arguments, Gineqnonlin, the gradient of
ineqnonlin(x), and Geqnonlin, the gradient of
eqnonlin(x). See Nonlinear Constraints for an explanation of how to “conditionalize” the
gradients for use in solvers that do not accept supplied gradients.
If nonlcon returns a vector ineqnonlin of
m components and x has length
n, where n is the length of
x0, then the gradient Gineqnonlin of
ineqnonlin(x) is an n-by-m
matrix, where Gineqnonlin(i,j) is the partial derivative of
ineqnonlin(j) with respect to x(i) (that is,
the jth column of Gineqnonlin is the gradient of
the jth inequality constraint ineqnonlin(j)).
Likewise, if eqnonlin has p components, the
gradient Geqnonlin of eqnonlin(x) is an
n-by-p matrix, where
Geqnonlin(i,j) is the partial derivative of
eqnonlin(j) with respect to x(i) (that is, the
jth column of Geqnonlin is the gradient of the
jth equality constraint eqnonlin(j)).
Note
Setting SpecifyConstraintGradient to true is
effective only when SpecifyObjectiveGradient is set to
true. Internally, the objective is folded into the
constraint, so the solver needs both gradients (objective and constraint) supplied
in order to avoid estimating a gradient.
See Passing Extra Parameters for an explanation of how to parameterize the
nonlinear constraint function nonlcon, if necessary.
Data Types: char | function_handle | string
Optimization options, specified as the output of
optimoptions or a structure as
optimset returns.
Some options apply to all algorithms, and others are relevant for particular algorithms. See Optimization Options Reference for detailed information.
Some options are absent from the
optimoptions display. These options appear in italics in the following
table. For details, see View Optimization Options.
| All Algorithms | |
| Choose between The The For more information on choosing the algorithm, see Choosing the Algorithm. |
| Diagnostics | Display diagnostic information
about the function to be minimized or solved. Choices are |
| DiffMaxChange | Maximum change in variables for
finite-difference gradients (a positive scalar). The default is |
| DiffMinChange | Minimum change in variables for
finite-difference gradients (a positive scalar). The default is |
| Level of display (see Iterative Display):
|
FiniteDifferenceStepSize | Scalar or vector step size factor for finite differences. When
you set
sign′(x) = sign(x) except sign′(0) = 1.
Central finite differences are
FiniteDifferenceStepSize expands to a vector. The default
is sqrt(eps) for forward finite differences, and eps^(1/3)
for central finite differences.
For |
FiniteDifferenceType | Finite differences, used to estimate gradients,
are either The algorithm is careful to obey bounds when estimating both types of finite differences. So, for example, it could take a backward, rather than a forward, difference to avoid evaluating at a point outside bounds. For |
| Termination tolerance on the function value, a nonnegative
scalar. The default is For
|
| FunValCheck | Check whether function values are
valid. |
| Maximum number of function evaluations allowed, a nonnegative
integer. The default is For |
| Maximum number of iterations allowed, a nonnegative integer. The
default is For |
OptimalityTolerance | Termination tolerance on the first-order optimality (a
nonnegative scalar). The default is Internally,
the For |
OutputFcn | Specify one or more user-defined functions that an optimization
function calls at each iteration. Pass a function handle or a cell array of function
handles. The default is none ( |
| Plots various measures of progress while the algorithm executes;
select from predefined plots or write your own. Pass a name, a function handle, or a
cell array of names or function handles. For custom plot functions, pass function
handles. The default is none (
Custom plot functions use the same syntax as output functions. See Output Functions for Optimization Toolbox and Output Function and Plot Function Syntax. For |
SpecifyObjectiveGradient | If For |
| Termination tolerance on For
|
| Typical |
UseParallel | When |
| Trust-Region-Reflective Algorithm | |
JacobianMultiplyFcn | Jacobian multiply function, specified as a function handle. For
large-scale structured problems, this function computes the Jacobian matrix product
W = jmfun(Jinfo,Y,flag) where
W = jmfun(Jinfo,Y,flag,xdata) where
The data [F,Jinfo] = fun(x)
% or [F,Jinfo] = fun(x,xdata)
In each case, Note
See Minimization with Dense Structured Hessian, Linear Equalities and Jacobian Multiply Function with Linear Least Squares for similar examples. For |
| JacobPattern | Sparsity pattern of the Jacobian
for finite differencing. Set Use If
the structure is unknown, do not set |
| MaxPCGIter | Maximum number of PCG (preconditioned
conjugate gradient) iterations, a positive scalar. The default is |
| PrecondBandWidth | Upper bandwidth of preconditioner
for PCG, a nonnegative integer. The default |
SubproblemAlgorithm | Determines how the iteration step
is calculated. The default, |
| TolPCG | Termination tolerance on the PCG
iteration, a positive scalar. The default is |
| Levenberg-Marquardt Algorithm | |
| InitDamping | Initial value of the Levenberg-Marquardt parameter, a
positive scalar. Default is |
| ScaleProblem |
|
UseCodegenSolver | Indication to use the version of the software that runs on
target hardware, specified as |
| Interior-Point Algorithm | |
BarrierParamUpdate | Specifies how
This option can affect the speed and convergence of the solver, but the effect is not easy to predict. |
ConstraintTolerance | Tolerance on the constraint violation, a nonnegative scalar. The default is
For
|
| InitBarrierParam | Initial barrier value, a positive scalar. Sometimes it might help to try a
value above the default |
SpecifyConstraintGradient | Gradient for nonlinear constraint functions defined by the user. When set to
the default, For |
SubproblemAlgorithm | Determines how the iteration step is calculated. The default,
For |
Example: options =
optimoptions('lsqnonlin','FiniteDifferenceType','central')
Problem structure, specified as a structure with the following fields:
| Field Name | Entry |
|---|---|
| Objective function |
| Initial point for
x |
| Matrix for linear inequality constraints |
| Vector for linear inequality constraints |
| Matrix for linear equality constraints |
| Vector for linear equality constraints |
lb | Vector of lower bounds |
ub | Vector of upper bounds |
| Nonlinear constraint function |
| 'lsqnonlin' |
| Options created with optimoptions |
You must supply at least the objective,
x0, solver, and
options fields in the problem
structure.
Data Types: struct
Output Arguments
Solution, returned as a real vector or real array. The size
of x is the same as the size of x0.
Typically, x is a local solution to the problem
when exitflag is positive. For information on
the quality of the solution, see When the Solver Succeeds.
Squared norm of the residual, returned as a nonnegative real.
resnorm is the squared 2-norm of the residual at
x: sum(fun(x).^2).
Value of objective function at solution, returned as an array. In general,
residual = fun(x).
Reason the solver stopped, returned as an integer.
| Function converged to a solution |
| Change in |
| Change in the residual is less than the specified tolerance. |
| Relative magnitude of search direction is smaller than the step tolerance. |
| Number of iterations exceeds |
| A plot function or output function stopped the solver. |
| No feasible point found. The bounds |
Information about the optimization process, returned as a structure with fields:
firstorderopt | Measure of first-order optimality |
iterations | Number of iterations taken |
funcCount | The number of function evaluations |
cgiterations | Total number of PCG iterations ( |
stepsize | Final displacement in |
constrviolation | Maximum of constraint functions
( |
bestfeasible | Best (lowest objective function) feasible point encountered
(
If no feasible point is found, the
The
|
algorithm | Optimization algorithm used |
message | Exit message |
Lagrange multipliers at the solution, returned as a structure with fields:
Jacobian at the solution, returned as a real matrix. jacobian(i,j) is
the partial derivative of fun(i) with respect to x(j) at
the solution x.
For problems with active constraints at the solution, jacobian is
not useful for estimating confidence intervals.
Limitations
The trust-region-reflective algorithm does not solve underdetermined systems; it requires that the number of equations, i.e., the row dimension of F, be at least as great as the number of variables. In the underdetermined case,
lsqnonlinuses the Levenberg-Marquardt algorithm.lsqnonlincan solve complex-valued problems directly. Note that constraints do not make sense for complex values, because complex numbers are not well-ordered; asking whether one complex value is greater or less than another complex value is nonsensical. For a complex problem with bound constraints, split the variables into real and imaginary parts. Do not use the'interior-point'algorithm with complex data. See Fit a Model to Complex-Valued Data.The preconditioner computation used in the preconditioned conjugate gradient part of the trust-region-reflective method forms JTJ (where J is the Jacobian matrix) before computing the preconditioner. Therefore, a row of J with many nonzeros, which results in a nearly dense product JTJ, can lead to a costly solution process for large problems.
If components of x have no upper (or lower) bounds,
lsqnonlinprefers that the corresponding components ofub(orlb) be set toinf(or-inffor lower bounds) as opposed to an arbitrary but very large positive (or negative for lower bounds) number.
You can use the trust-region reflective algorithm in lsqnonlin, lsqcurvefit,
and fsolve with small- to medium-scale
problems without computing the Jacobian in fun or
providing the Jacobian sparsity pattern. (This also applies to using fmincon or fminunc without
computing the Hessian or supplying the Hessian sparsity pattern.)
How small is small- to medium-scale? No absolute answer is available,
as it depends on the amount of virtual memory in your computer system
configuration.
Suppose your problem has m equations and n unknowns.
If the command J = sparse(ones(m,n)) causes
an Out of memory error on your machine,
then this is certainly too large a problem. If it does not result
in an error, the problem might still be too large. You can find out
only by running it and seeing if MATLAB runs within the amount
of virtual memory available on your system.
More About
The next few items list the possible enhanced exit messages from
lsqnonlin. Enhanced exit messages give a link for more
information as the first sentence of the message.
The solver located a point that seems to be a local minimum of the sum of squares, since the first-order optimality measure is close to 0.
For suggestions on how to proceed, see When the Solver Succeeds.
The solver may have reached a local minimum but cannot be
certain because the first-order optimality measure is not less than the
OptimalityTolerance
tolerance
(1e-4*OptimalityTolerance for the Levenberg-Marquardt
algorithm).
For suggestions on how to proceed, see Local Minimum Possible.
The initial point seems to be a local minimum of the sum of squares, because the first-order optimality measure is close to 0.
For suggestions on how to proceed, see Final Point Equals Initial Point.
The next few items contain definitions for terms in the lsqnonlin exit messages.
A local minimum of a function is a point where the function value is smaller than at nearby points, but possibly greater than at a distant point.
A global minimum is a point where the function value is smaller than at all other feasible points.
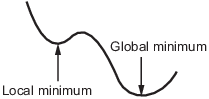
Solvers try to find a local minimum. The result can be a global minimum. For more information, see Local vs. Global Optima.
For unconstrained problems, the first-order optimality measure is the maximum of the absolute value of the components of the gradient vector (also known as the infinity norm of the gradient). This should be zero at a minimizing point.
For problems with bounds, the first-order optimality measure is the maximum over i of |vi*gi|. Here gi is the ith component of the gradient, x is the current point, and
If xi is at a bound, vi is zero. If xi is not at a bound, then at a minimizing point the gradient gi should be zero. Therefore the first-order optimality measure should be zero at a minimizing point.
For more information, see First-Order Optimality Measure.
Generally, a tolerance is a threshold which, if crossed, stops the iterations of a solver. For more information on tolerances, see Tolerances and Stopping Criteria.
The tolerance called OptimalityTolerance relates to the first-order optimality measure. Iterations end when
the first-order optimality measure is less than
OptimalityTolerance.
The function tolerance called
FunctionTolerance relates to the size of the latest change in
objective function value.

The gradient vector is the gradient of the sum of squares. For unconstrained problems, the gradient size is the maximum of the absolute value of the components of the gradient vector (also known as the infinity norm of the gradient). This should be zero at a minimizing point.
For problems with bounds, the gradient size is the maximum over i of |vi*gi|. Here gi is the ith component of the gradient, x is the current point, and
If xi is at a bound, vi is zero. If xi is not at a bound, then at a minimizing point the gradient gi should be zero. Therefore the gradient size should be zero at a minimizing point.
For more information, see First-Order Optimality Measure.
The size of the current step is the magnitude of the change in location of the current point in the final iteration.
For more information, see Tolerances and Stopping Criteria.
The Levenberg-Marquardt regularization parameter is related to the inverse of a trust-region radius. It becomes large when the sum of squares of function values is not close to a quadratic model. For more information, see Levenberg-Marquardt Method.
Solvers estimate the Jacobian of your objective vector function by taking finite
differences. A finite difference calculation stepped outside the region where the
objective function is well-defined, returning Inf,
NaN, or a complex result.
For more information about how solvers use the Jacobian J, see Levenberg-Marquardt Method. For suggestions on how to proceed, see 6. Provide Gradient or Jacobian.
Algorithms
The Levenberg-Marquardt and trust-region-reflective methods are based on the nonlinear
least-squares algorithms also used in fsolve.
The default trust-region-reflective algorithm is a subspace trust-region method and is based on the interior-reflective Newton method described in [1] and [2]. Each iteration involves the approximate solution of a large linear system using the method of preconditioned conjugate gradients (PCG). See Trust-Region-Reflective Least Squares.
The Levenberg-Marquardt method is described in references [4], [5], and [6]. See Levenberg-Marquardt Method.
The 'interior-point' algorithm uses the fmincon
'interior-point' algorithm with some modifications. For details, see
Modified fmincon Algorithm for Constrained Least Squares.
Alternative Functionality
App
The Optimize Live Editor task provides a visual interface for lsqnonlin.
References
[1] Coleman, T.F. and Y. Li. “An Interior, Trust Region Approach for Nonlinear Minimization Subject to Bounds.” SIAM Journal on Optimization, Vol. 6, 1996, pp. 418–445.
[2] Coleman, T.F. and Y. Li. “On the Convergence of Reflective Newton Methods for Large-Scale Nonlinear Minimization Subject to Bounds.” Mathematical Programming, Vol. 67, Number 2, 1994, pp. 189–224.
[3] Dennis, J. E. Jr. “Nonlinear Least-Squares.” State of the Art in Numerical Analysis, ed. D. Jacobs, Academic Press, pp. 269–312.
[4] Levenberg, K. “A Method for the Solution of Certain Problems in Least-Squares.” Quarterly Applied Mathematics 2, 1944, pp. 164–168.
[5] Marquardt, D. “An Algorithm for Least-squares Estimation of Nonlinear Parameters.” SIAM Journal Applied Mathematics, Vol. 11, 1963, pp. 431–441.
[6] Moré, J. J. “The Levenberg-Marquardt Algorithm: Implementation and Theory.” Numerical Analysis, ed. G. A. Watson, Lecture Notes in Mathematics 630, Springer Verlag, 1977, pp. 105–116.
[7] Moré, J. J., B. S. Garbow, and K. E. Hillstrom. User Guide for MINPACK 1. Argonne National Laboratory, Rept. ANL–80–74, 1980.
[8] Powell, M. J. D. “A Fortran Subroutine for Solving Systems of Nonlinear Algebraic Equations.” Numerical Methods for Nonlinear Algebraic Equations, P. Rabinowitz, ed., Ch.7, 1970.
Extended Capabilities
lsqcurvefitandlsqnonlinsupport code generation using either thecodegen(MATLAB Coder) function or the MATLAB Coder™ app. You must have a MATLAB Coder license to generate code.The target hardware must support standard double-precision floating-point computations. You cannot generate code for single-precision or fixed-point computations.
Code generation targets do not use the same math kernel libraries as MATLAB solvers. Therefore, code generation solutions can vary from solver solutions, especially for poorly conditioned problems.
To test your code in MATLAB before generating code, set the
UseCodegenSolveroption totrue. That way, the solver uses the same code that code generation creates.All code for generation must be MATLAB code. In particular, you cannot use a custom black-box function as an objective function for
lsqcurvefitorlsqnonlin. You can usecoder.cevalto evaluate a custom function coded in C or C++. However, the custom function must be called in a MATLAB function.Code generation for
lsqcurvefitandlsqnonlincurrently does not support linear or nonlinear constraints.Code generation does not support complex-valued data.
lsqcurvefitandlsqnonlindo not support theproblemargument for code generation.[x,fval] = lsqnonlin(problem) % Not supportedYou must specify the objective function by using function handles, not strings or character names.
x = lsqnonlin(@fun,x0,lb,ub,options) % Supported % Not supported: lsqnonlin('fun',...) or lsqnonlin("fun",...)
All input matrices
lbandubmust be full, not sparse. You can convert sparse matrices to full by using thefullfunction.The
lbandubarguments must have the same number of entries as thex0argument or must be empty[].If your target hardware does not support infinite bounds, use
optim.coder.infbound.For advanced code optimization involving embedded processors, you also need an Embedded Coder® license.
You must include options for
lsqcurvefitorlsqnonlinand specify them usingoptimoptions. The options must include theAlgorithmoption, set to"levenberg-marquardt".options = optimoptions("lsqnonlin",Algorithm="levenberg-marquardt"); [x,fval,exitflag] = lsqnonlin(fun,x0,lb,ub,options);
Code generation supports these options:
Algorithm— Must be"levenberg-marquardt"FiniteDifferenceStepSizeFiniteDifferenceTypeFunctionToleranceMaxFunctionEvaluationsMaxIterationsSpecifyObjectiveGradientStepToleranceTypicalXUseCodegenSolver
Generated code has limited error checking for options. The recommended way to update an option is to use
optimoptions, not dot notation.opts = optimoptions("lsqnonlin",Algorithm="levenberg-marquardt"); opts = optimoptions(opts,MaxIterations=1e4); % Recommended opts.MaxIterations = 1e4; % Not recommended
Do not load options from a file. Doing so can cause code generation to fail. Instead, create options in your code.
Usually, if you specify an option that is not supported, the option is silently ignored during code generation. However, if you specify a plot function or output function by using dot notation, code generation can issue an error. For reliability, specify only supported options.
Because output functions and plot functions are not supported, solvers do not return the exit flag –1.
For an example, see Generate Code for lsqcurvefit or lsqnonlin.
To run in parallel, set the 'UseParallel' option to true.
options = optimoptions('solvername','UseParallel',true)
For more information, see Using Parallel Computing in Optimization Toolbox.
Version History
Introduced before R2006aSet the new UseCodegenSolver option to true to have lsqnonlin use the same version of the software that code generation creates. This option allows you to check the behavior of the solver before you generate code or deploy the code to hardware. You can include the option when you generate code; the option has no effect in code generation, but leaving the option in saves you the step of removing it. Even though the generated code is identical to the MATLAB code, results can differ slightly because linked math libraries can differ.
The CheckGradients option has been removed. To check whether your code computes gradients correctly, use the checkGradients function. For example,
function [f,g] = rosen(x) f = 100*(x(1) - x(2)^2)^2 + (1 - x(2))^2; if nargout > 1 g(1) = 200*(x(1) - x(2)^2); g(2) = -400*x(2)*(x(1) - x(2)^2) - 2*(1 - x(2)); end end % Before using the rosen function, % you can check that the gradient is correct at a point x0 = [2,4]; assert(checkGradients(@rosen,x0))
The syntax for the JacobianMultiplyFcn option is
W = jmfun(Jinfo, Y, flag)
The Jinfo data, which MATLAB passes to your function jmfun, can now be of any data
type. For example, you can now have Jinfo be a structure. In previous
releases, Jinfo had to be a standard double array.
The Jinfo data is the second output of your objective
function:
[F,Jinfo] = myfun(x)
The CheckGradients option will be removed in a future release. To check the first derivatives of objective functions or nonlinear constraint functions, use the checkGradients function.
lsqnonlin gains support for both linear and nonlinear constraints. To enable constraint satisfaction, the solver uses the "interior-point" algorithm from fmincon.
If you specify constraints but do not specify an algorithm, the solver automatically switches to the
"interior-point"algorithm.If you specify constraints and an algorithm, you must specify the
"interior-point"algorithm.
For algorithm details, see Modified fmincon Algorithm for Constrained Least Squares. For an example, see Compare lsqnonlin and fmincon for Constrained Nonlinear Least Squares.
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)