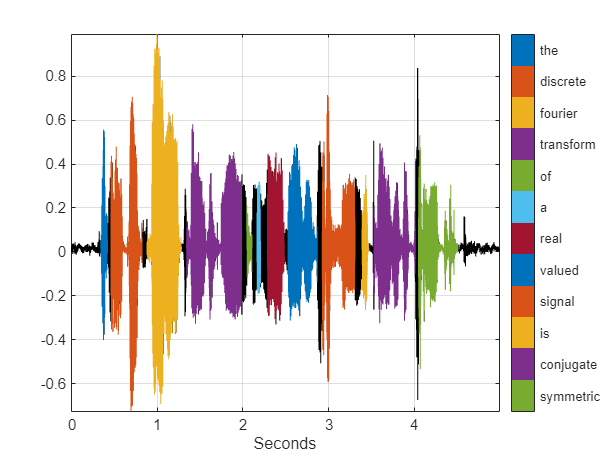
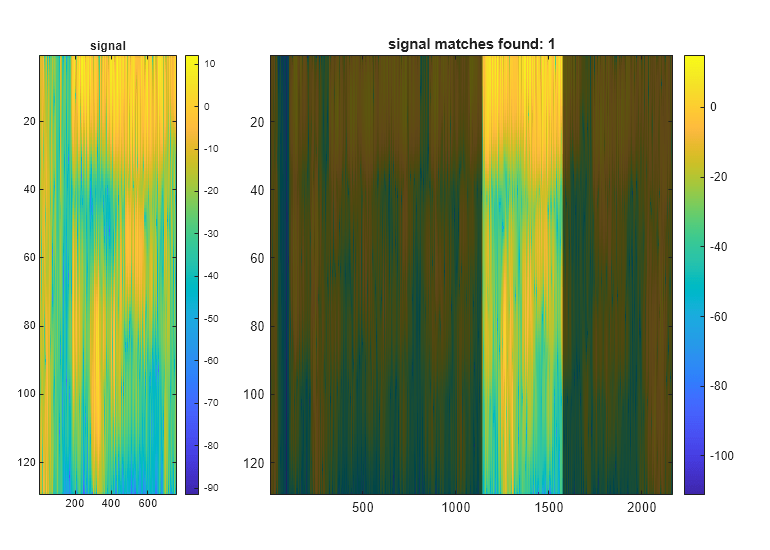
Descriptive Statistics
Use findpeaks to locate the local maxima of a signal and sort the peaks by height, width, or prominence. Determine the crest factor of a signal using the peak2rms function and compute common descriptive statistics like maxima, minima, standard deviations, and RMS levels. Search for signals of interest in larger data sets and align signals in time. Locate points where a signal changes abruptly or drifts beyond a target range. Label signals for analysis or machine and deep learning applications.
Apps
| Signal Analyzer | Visualize and compare multiple signals and spectra |
| Signal Labeler | Label signal attributes, regions, and points of interest |
| Signal Feature Extractor | Extract and analyze signal features (Since R2025a) |
Functions
Topics
- Use Signal Feature Extractor App
Extract and rank features from signals interactively. Prepare signal datasets for classification tasks.
- RMS Value of Periodic Waveforms
Find the root mean square value of a sine wave, a square wave, and a rectangular pulse train.
- Find Peaks in Data
Locate the local maxima in a set of data and determine if those peaks occur periodically.
- Prominence
The prominence of a peak measures how much the peak stands out due to its intrinsic height and its location relative to other peaks.
- Human Activity Recognition Simulink Model for Smartphone Deployment (Statistics and Machine Learning Toolbox)
Generate code from a classification Simulink® model prepared for deployment to a smartphone.
- Choose an App to Label Ground Truth Data
Decide which app to use to label ground truth data: Image Labeler, Video Labeler, Ground Truth Labeler, Lidar Labeler, Signal Labeler, or Medical Image Labeler.