clusterDBSCAN
Density-based algorithm for clustering data
Description
clusterDBSCAN clusters data points belonging to a
P-dimensional feature space using the density-based spatial clustering of
applications with noise (DBSCAN) algorithm. The clustering algorithm assigns points that are
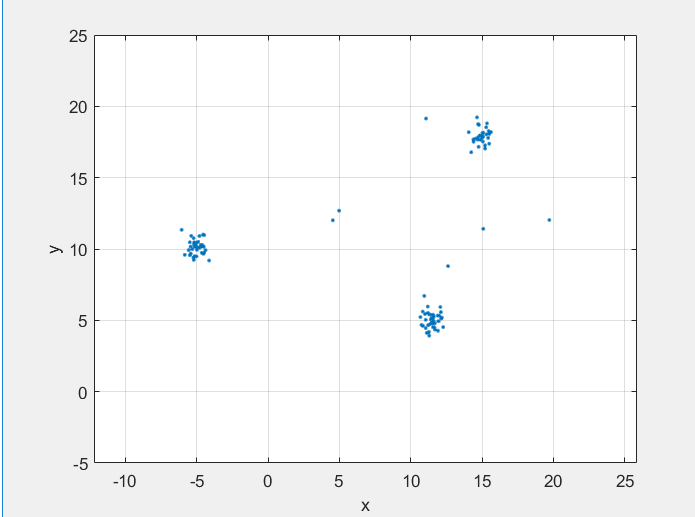
close to each other in feature space to a single cluster. For example, a radar system can
return multiple detections of an extended target that are closely spaced in range, angle, and
Doppler. clusterDBSCAN assigns these detections to a single detection.
The DBSCAN algorithm assumes that clusters are dense regions in data space separated by regions of lower density and that all dense regions have similar densities.
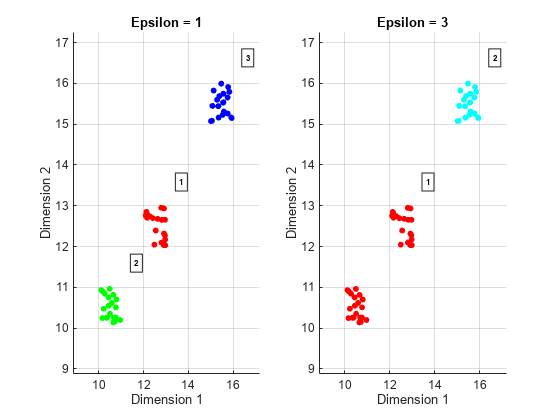
To measure density at a point, the algorithm counts the number of data points in a neighborhood of the point. A neighborhood is a P-dimensional ellipse (hyperellipse) in the feature space. The radii of the ellipse are defined by the P-vector ε. ε can be a scalar, in which case, the hyperellipse becomes a hypersphere. Distances between points in feature space are calculated using the Euclidean distance metric. The neighborhood is called an ε-neighborhood. The value of ε is defined by the
Epsilonproperty.Epsiloncan either be a scalar or P-vector:A vector is used when different dimensions in feature space have different units.
A scalar applies the same value to all dimensions.
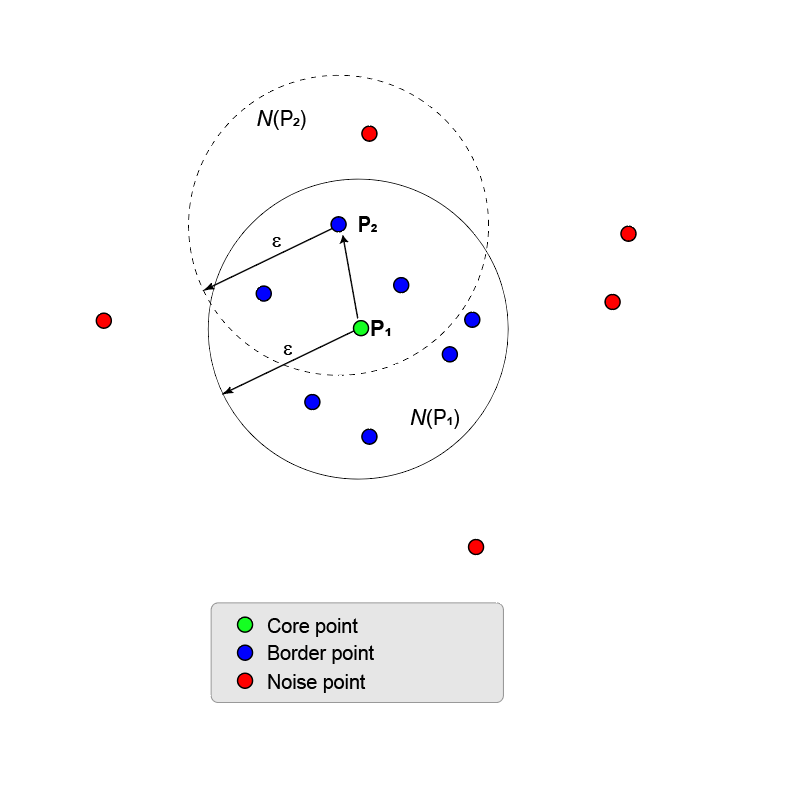
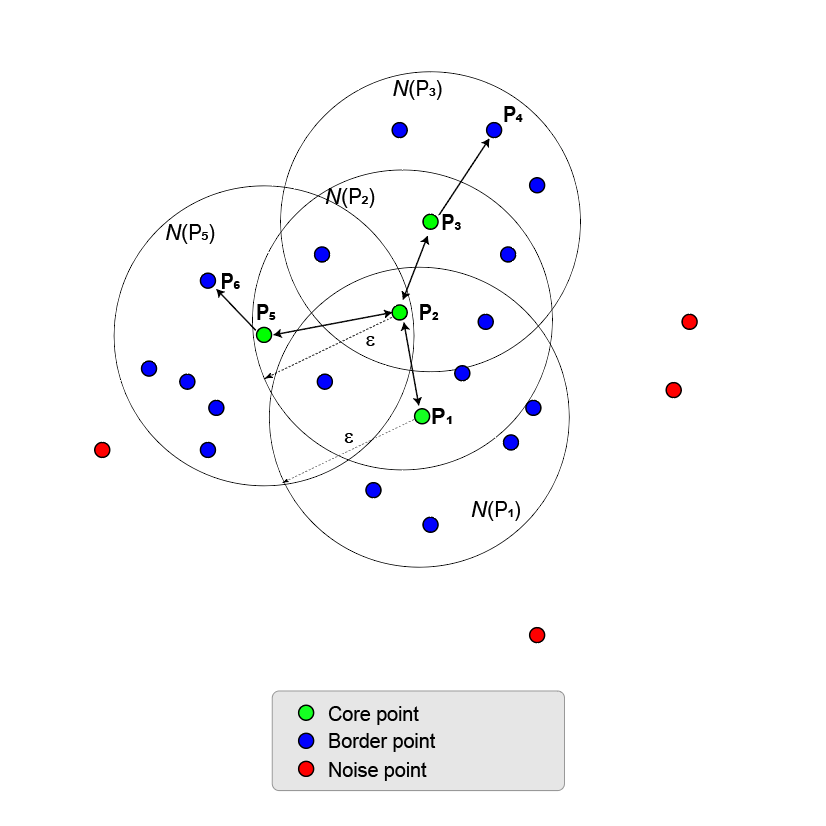
Clustering starts by finding all core points. If a point has a sufficient number of points in its ε-neighborhood, the point is called a core point. The minimum number of points required for a point to become a core point is set by the
MinNumPointsproperty.The remaining points in the ε-neighborhood of a core point can be core points themselves. If not, they are border points. All points in the ε-neighborhood are called directly density reachable from the core point.
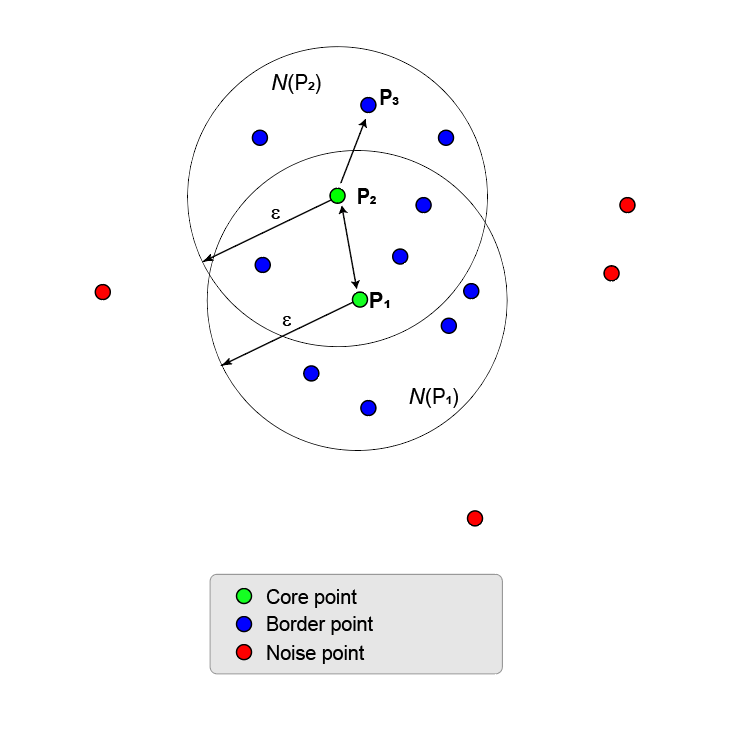
If the ε-neighborhood of a core point contains other core points, the points in the ε-neighborhoods of all the core points merge together to form a union of ε-neighborhoods. This process continues until no more core points can be added.
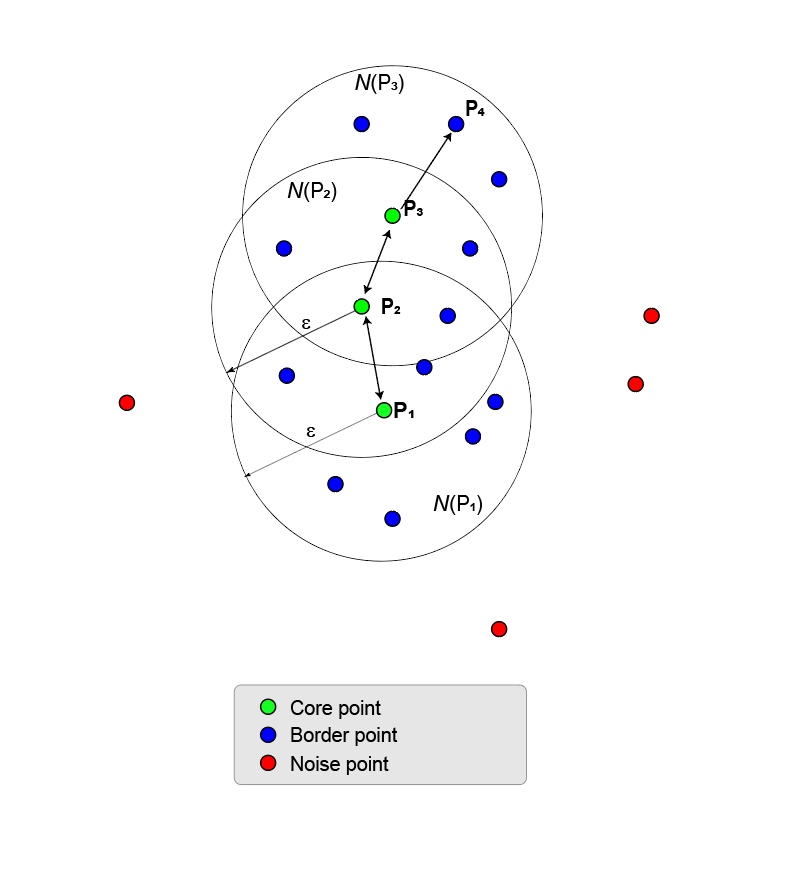
All points in the union of ε-neighborhoods are density reachable from the first core point. In fact, all points in the union are density reachable from all core points in the union.
All points in the union of ε-neighborhoods are also termed density connected even though border points are not necessarily reachable from each other. A cluster is a maximal set of density-connected points and can have an arbitrary shape.
Points that are not core or border points are noise points. They do not belong to any cluster.
The
clusterDBSCANobject can estimate ε using a k-nearest neighbor search, or you can specify values. To let the object estimate ε, set theEpsilonSourceproperty to'Auto'.The
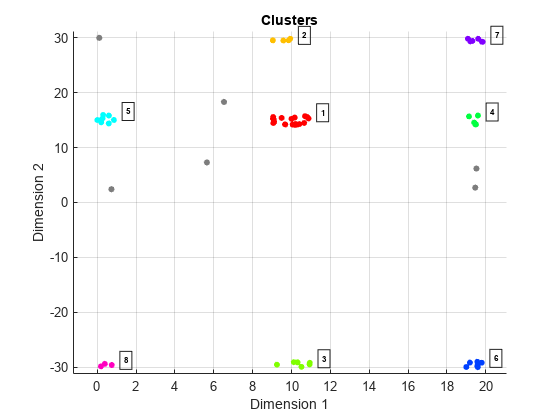
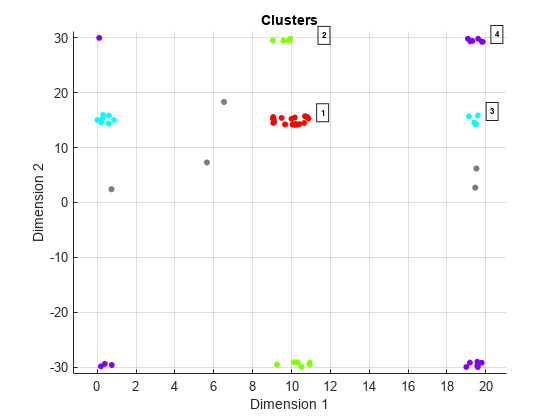
clusterDBSCANobject can disambiguate data containing ambiguities. Range and Doppler are examples of possibly ambiguous data. SetEnableDisambiguationproperty totrueto disambiguate data.
To cluster detections:
Create the
clusterDBSCANobject and set its properties.Call the object with arguments, as if it were a function.
To learn more about how System objects work, see What Are System Objects?
Creation
Description
clusterer = clusterDBSCANclusterDBSCAN object, clusterer, with default
property values.
clusterer = clusterDBSCAN(Name,Value)clusterDBSCAN object, clusterer, with each
specified Property
Name set to the corresponding Value. You can
specify additional pairs of arguments in any order as
Name1=Value1,...,NameN=ValueN. Any unspecified
properties take default values. For example,
clusterer = clusterDBSCAN(MinNumPoints=3,Epsilon=2, ...
EnableDisambiguation=true,AmbiguousDimension=[1 2]);clusterer object with the EnableDisambiguation
property set to true and the AmbiguousDimension set to
[1,2].Properties
Usage
Syntax
Description
[
also returns an alternate set of cluster IDs, idx,clusterids] = clusterer(X)clusterids, for use in
the phased.RangeEstimator and phased.DopplerEstimator objects. clusterids assigns a
unique ID to each noise point.
[___] = clusterer(
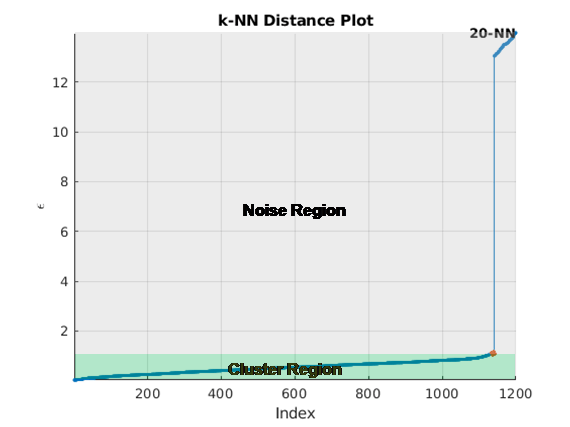
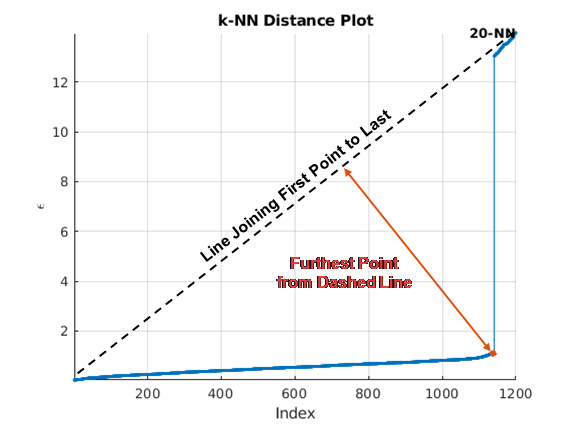
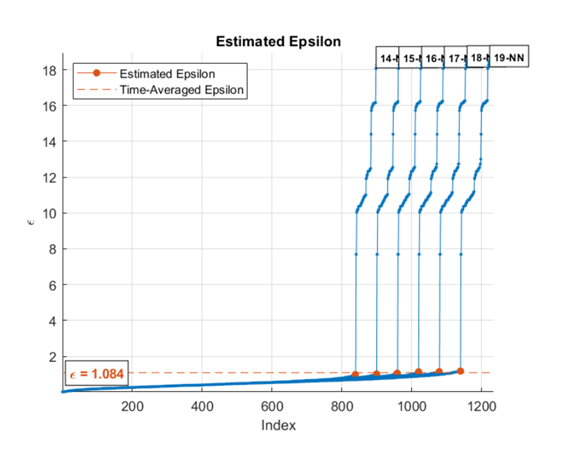
automatically estimates epsilon from the input data matrix, X,update)X, when
update is set to true. The estimation uses a
k-NN search to create a set of search curves. For more information,
see Estimate Epsilon. The estimate is an
average of the L most recent Epsilon values where L
is specified in EpsilonHistoryLength
To enable this syntax, set the EpsilonSource property to
'Auto', optionally set the MaxNumPoints
property, and also optionally set the EpsilonHistoryLength
property.
Input Arguments
Output Arguments
Object Functions
To use an object function, specify the
System object™ as the first input argument. For
example, to release system resources of a System object named obj, use
this syntax:
release(obj)
Examples
Algorithms
References
[1] Ester M., Kriegel H.-P., Sander J., and Xu X. "A Density-Based Algorithm for Discovering Clusters in Large Spatial Databases with Noise". Proc. 2nd Int. Conf. on Knowledge Discovery and Data Mining, Portland, OR, AAAI Press, 1996, pp. 226-231.
[2] Erich Schubert, Jörg Sander, Martin Ester, Hans-Peter Kriegel, and Xiaowei Xu. 2017. "DBSCAN Revisited, Revisited: Why and How You Should (Still) Use DBSCAN". ACM Trans. Database Syst. 42, 3, Article 19 (July 2017), 21 pages.
[3] Dominik Kellner, Jens Klappstein and Klaus Dietmayer, "Grid-Based DBSCAN for Clustering Extended Objects in Radar Data", 2012 IEEE Intelligent Vehicles Symposium.
[4] Thomas Wagner, Reinhard Feger, and Andreas Stelzer, "A Fast Grid-Based Clustering Algorithm for Range/Doppler/DoA Measurements", Proceedings of the 13th European Radar Conference.
[5] Mihael Ankerst, Markus M. Breunig, Hans-Peter Kriegel, Jörg Sander, "OPTICS: Ordering Points To Identify the Clustering Structure", Proc. ACM SIGMOD’99 Int. Conf. on Management of Data, Philadelphia PA, 1999.
Extended Capabilities
Version History
Introduced in R2021a